I’ve been playing with Steffen Müller’s Dumbbench module, which applies some statistical analysis to benchmarking. He explains it all in a couple of blog posts:
The benchmarking part is interesting, but the more interesting part is that Dumbbench can make pictures, unlike the boring, old, ASCII-reporting Benchmark.
Steffen used an interface to ROOT, a huge toolkit out of CERN. I couldn’t compile it on my Mac, so I started playing with R to make my plots. Instead of using ROOT, I added a feature to write the data to a table so I could feed that table to Rscript:
$ dumbbench --code='++$i' --code='$i++' --code='$i+=1' --table=code.dat --maxiter 10000 $ Rscript --vanilla --slave r/boxplot.r code.dat
From that I get an impressive picture that looks as good as the one from ROOT:
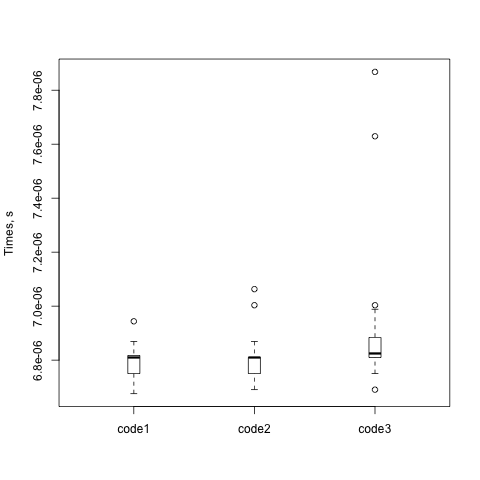
I don’t want to supply code on the command line, though. Dumbbench has three sources (or, instances). There’s that --code switch and there’s a final command line argument. The module can take subroutines too, so I added a --package switch. I can put my code to benchmark in a file that I can save and edit and reüse.
$ dumbbench --package=SomePackage --table=some_package.dat --maxiter 10000
There are still some things I want to do. I’d love to figure out how R’s read_table can take its data from standard input so I can get rid of the intermediate file and pipe directly to R.
More interesting than that, however, is fancy stuff I might be able to do in that package. Every subroutine should accomplish the same task, and perhaps return the same thing. The package would be able to automatically test each of the subroutines to verify that. I haven’t thought too much about that yet.